Protein Design and Engineering
Outline of Research and Education
At the Laboratory of Protein Design and Engineering, we investigate the fundamental relationship between protein structure and biological function. Utilizing an interdisciplinary approach that combines structural biology, biochemistry, and biophysics, we decode the molecular mechanisms underlying enzymatic catalysis, intracellular signaling, and molecular recognition. We believe that a deep structural understanding is the ultimate key to engineering superior proteins.
Guided by our core philosophy: "Understand – Enhance – Design", we push the boundaries of protein engineering by integrating rapidly evolving machine learning (ML) technologies. Our current research focuses on the mechanistic control of non-template polymerases for biodegradable plastic synthesis, alongside the development of enzymes capable of upcycling petroleum-based plastics. Through rigorous basic research, we aim to pioneer sustainable material development and shape the future of biotechnology for a circular economy.
In terms of education, we provide comprehensive training in laboratory safety and fundamental techniques. We foster an environment where creativity and free discussion are encouraged to spark innovation. Furthermore, we emphasize English as the primary medium of communication and presentation to prepare our students for effective global scientific discourse.
Major Research Topics
Elucidation of Non-Template-Based Polymerization by PHA synthase
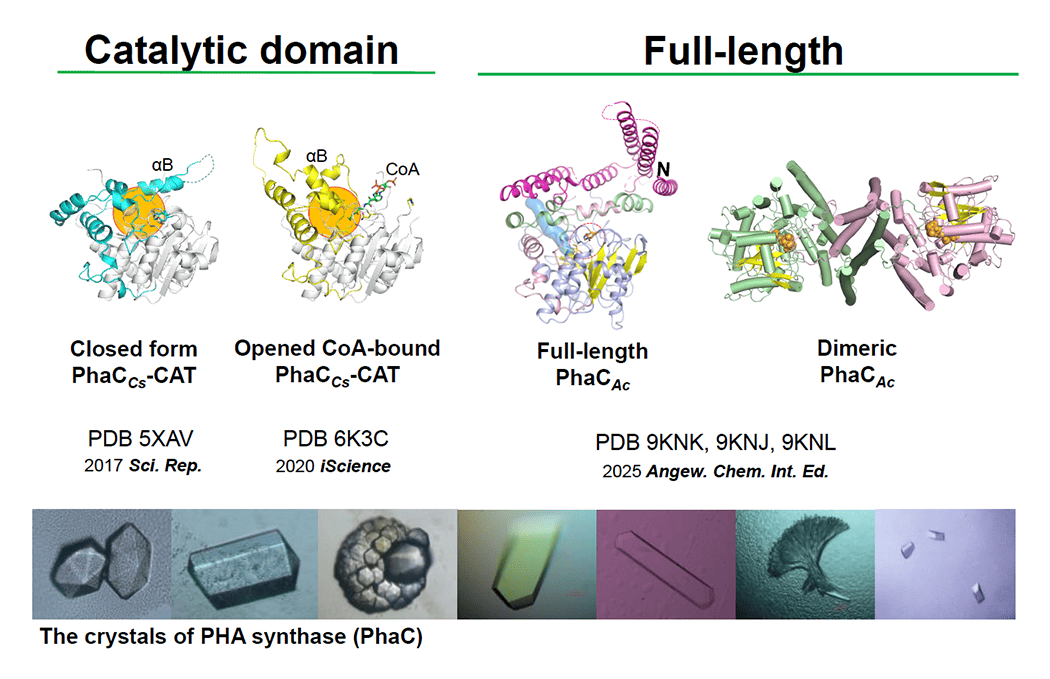
Polyhydroxyalkanoates (PHAs) are biodegradable polyesters produced by many microorganisms and are promising alternatives to petrochemical plastics. PHAs are classified into short-chain-length (scl-PHA), which are rigid and brittle; medium-chain-length (mcl-PHA), which are soft and sticky; and scl–mcl copolymers, which are elastic and comparable to commodity plastics. PHA synthase (PhaC) is the key enzyme that determines the type and composition of PHA produced. However, most natural PhaCs mainly synthesize brittle scl-PHAs. Therefore, increasing the production of flexible scl–mcl copolymers, such as poly(3-hydroxybutyrate-co-3-hydroxyhexanoate), is important for expanding practical applications.
Despite its central role, the catalytic mechanism of PhaC remains unclear. Key questions include whether it follows a ping-pong or processive mechanism, how polymer chain termination occurs, and how monomer incorporation is regulated. Answering these questions will improve our understanding the fundamental principles of template-independent polymer synthesis.
This project aims to clarify these mechanisms using biochemical and structural approaches, enabling the rational engineering of customized PhaC variants for advanced biodegradable polymer production. Our work combines molecular cloning, X-ray crystallography, cryo-electron microscopy, computational structure prediction, and comprehensive biochemical and biophysical analyses. We have already solved multiple three-dimensional structures of PhaC from different sources and will further investigate key mechanistic intermediates (Fig. 1).
Protein Engineering of Enzymes for Sustainable Development
Proteins are essential workhorses in both industrial processes and biological systems. However, most naturally occurring enzymes do not meet the performance requirements needed for industrial or therapeutic applications. To overcome these limitations, we engineer enzymes to improve thermal stability, catalytic efficiency, and substrate specificity using structure-guided protein engineering. Our approach combines computational analysis with strong experimental validation. We identify key residues involved in catalysis and substrate binding, introduce targeted mutations based on structural insights and sequence alignments, and then express, purify, and evaluate the engineered variants. Through iterative testing and refinement, we optimize protein performance using rational design and directed evolution. Our current research focuses on enzymes for biodegradable plastic synthesis and petrochemical plastic degradation, contributing to sustainable development and SDG 14 (Life Below Water) (Fig. 2). We also work on enhancing protein–ligand binding affinity for therapeutic applications. Our laboratory is equipped with advanced analytical tools to study molecular interactions, including isothermal titration calorimetry (ITC) for protein–ligand binding, biolayer interference (BLI) for protein–protein interactions, and a Varioskan high-throughput screening system with FRET and AlphaScreen capabilities for inhibitor discovery. We welcome industrial and academic collaborations, as well as new research ideas from students.
Design of FusionTags for Cryo-EM structure determination
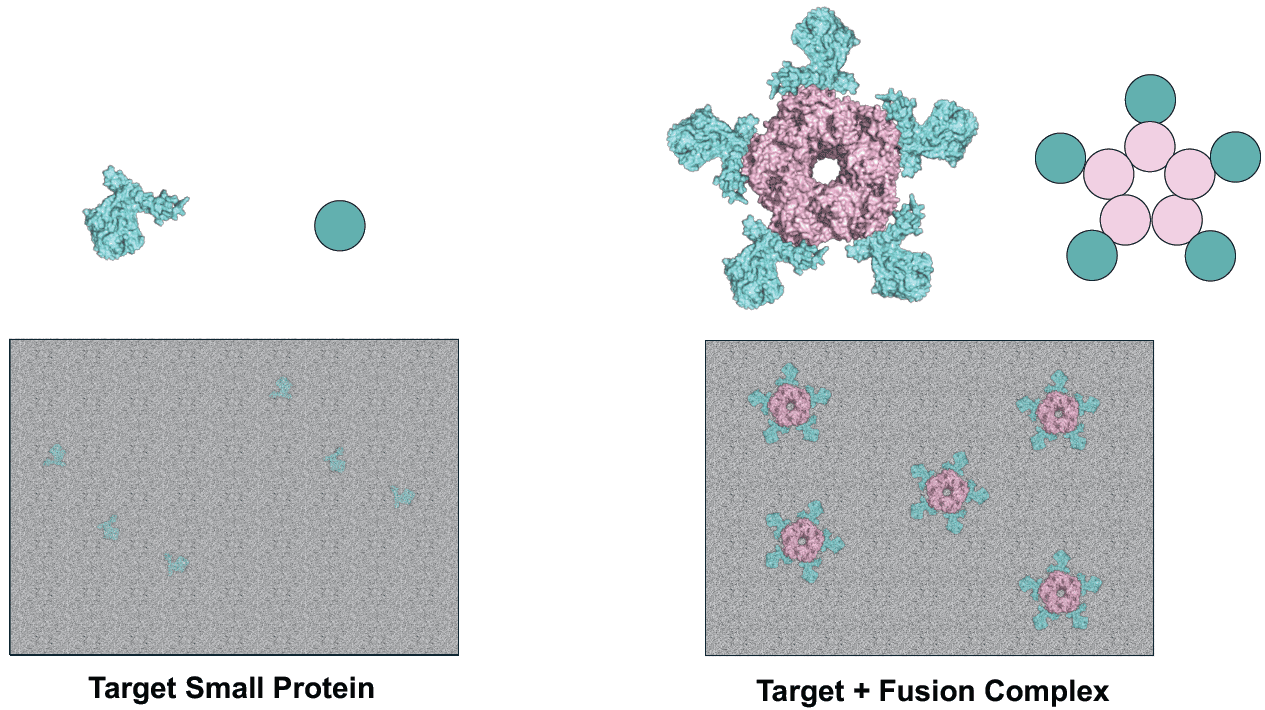
Recent advances in cryo-electron microscopy (cryo-EM) have enabled near-atomic resolution structure determination for a wide array of enzymatic proteins. However, the structural analysis of small (<50 kDa) or flexible proteins remains a significant challenge due to inherent limitations in molecular weight and conformational heterogeneity, which hinder accurate particle alignment. To address these issues, our research focuses on the rational design of stable fusion scaffolds to enhance structural rigidity and facilitate precise orientation during cryo-EM data processing. Our strategy involves both the optimization of existing structural tags and the de novo design of novel scaffolds using machine-learning tools. By developing these specialized rigid tags, we aim to extend the applicability of cryo-EM to a broader range of recalcitrant protein targets. This multidisciplinary project integrates computational protein design, expression, purification, and structural validation, conducted in collaboration with the Indonesian National Research and Innovation Agency (BRIN) under the Japan-Indonesia JST NEXUS project on Biomonotsukuri (Bioproduction) (Fig. 3).

References
- Chek MF et al., International Journal of Biological Macromolecules, 149745, 2026
- Laohawutthichai P et al., Journal of Molecular Biology, 169329, 2025
- Chek MF* et al., Angewandte Chemie International Edition, 64, e202504626, 2025
- Qin TX et al., Cancer Letters, 628, 2025
- Sim PF†, Chek MF†, Nguyen NTH†et al., The Journal of Biochemistry, 175, 2024
- Tan HT et al., Polymer Degradation and Stability, 206, 2022
- Neoh SZ†, Chek MF† et al., Current Research in Biotechnology, 4, 2022
- Kim SY†, Mori T†, Chek MF† et al., Scientific Reports, 11(1), 2120, 2021
- Chek MF et al., iScience, 23(5), 101084, 2020
- Chek MF et al., Applied Microbiology and Biotechnology, 103, 2018
- Chek MF et al., Scientific Reports, 7, 5213, 2017
† Equal contribution; * Corresponding author


