Microbial Interaction

- Associate Professor
- WATANABE Daisuke

- Assistant Professor
- AKASAKA Naoki

- Labs HP
- https://bsw3.naist.jp/microbial_interaction/en/top/
Outline of Research and Education
We aim to explore how microorganisms closely related to humans (such as yeast, lactic acid bacteria, and koji fungus) behave and interact with other organisms and environmental factors to build complex ecosystems. Our research focuses on understanding the diversity in the microbial world at the molecular, metabolic, and cellular levels. Our achievement will also contribute to modern biotechnology in food and health science (Fig. 1).
Major Research Topics
Decoding yeast traits to design fermentation
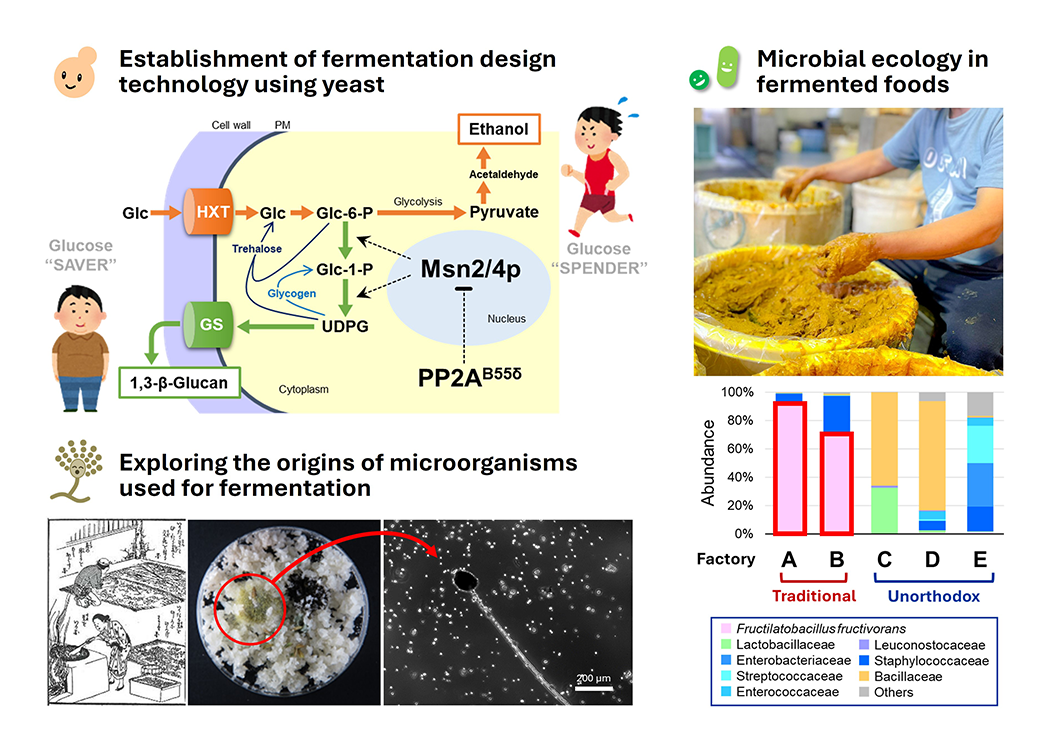
Alcohol fermentation by yeast is one of the microbial functions most closely connected to human life, supporting the production of alcoholic beverages and fermented foods. Although our understanding of yeast metabolic pathways and related genes has advanced considerably, it is still not easy to manipulate fermentative capacity per cell as desired. We focus on sake yeast, a unique microbial resource from Japan, and investigate the molecular basis underlying its high fermentation performance. We have shown that characteristic mutations found in sake yeast contribute to its high ethanol productivity, and have successfully applied this knowledge to improve the fermentation performance of other industrial yeasts and to develop yeast strains suitable for low-alcohol sake making. Moving forward, we aim not only to understand fermentation performance but also the traits of yeast related to flavor formation, and to establish a unique “fermentation design technology” that enables us to tailor fermentation for specific purposes (Fig. 2, upper left).
Unraveling the microbial ecosystems of traditional fermented foods
Traditional fermented foods around the world are supported not by a single microorganism but by complex ecosystems in which diverse microbes coexist and interact. However, many questions remain as to how stable fermentation is maintained through microbial interactions. We study traditional fermented foods such as sake and narazuke to clarify the microorganisms involved and the relationships among them. Our recent work revealed interactions between lactic acid bacteria and yeast in kimoto-style sake brewing and showed that lactic acid bacteria can influence alcoholic fermentation by yeast. We have also identified unusual lactic acid bacteria associated with narazuke and scientifically demonstrated that narazuke is a fermented food sustained by microbial activity. Through these studies, we aim to understand the principles underlying traditional fermented food production, rediscover their value, stabilize their manufacture, and generate knowledge that leads to new developments in the field (Fig. 2, right).
Exploring the origins of fermented foods and fermentation microorganisms
In modern fermentation industries, microorganisms selected for specific purposes are used in production. However, it remains unclear how people first began to harness fermentation before microorganisms were even known to exist. Because many fermented foods are produced from plant materials, plant-associated microbes may have played key roles in the origins of fermentation. Using old documents preserved in temples and records of traditional manufacturing methods as clues, we seek to uncover the roots of lost fermented foods and fermentation microorganisms through modern science. In particular, we are investigating koji fungus, which is indispensable for the production of sake, miso, and soy sauce, by reproducing natural inoculation processes using rice straw and by elucidating the mechanisms by which this fungus adapts to plant surfaces. Recently, we demonstrated that some microorganisms, including koji fungus, can grow by utilizing components of plant surface layers, bringing us closer to understanding how they establish themselves as plant-associated microbes. These studies are expected not only to clarify the origins of fermentation culture but also to contribute to the revival of traditional fermented foods and the development of new fermentation technologies using natural microorganisms (Fig. 2, lower left).
Discovering new health-promoting functions of fermented foods
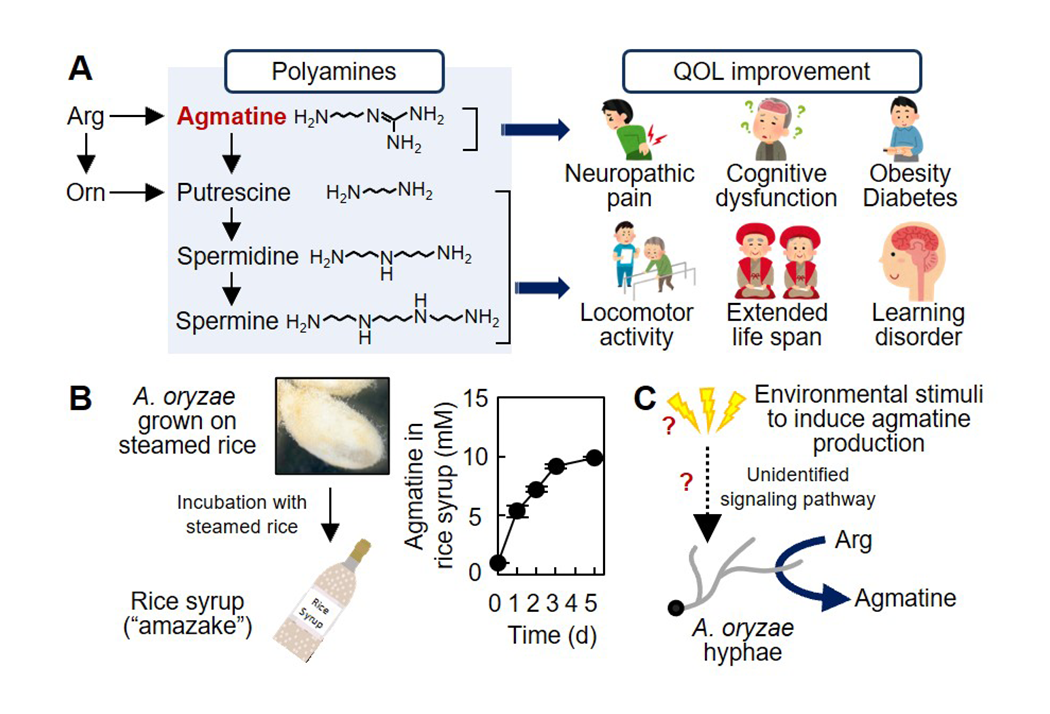
A variety of traditional fermented foods have been produced since ancient times in Japan, and have been contributing to Japanese healthy life span. However, why the fermented foods are beneficial for our health remains scientifically well understood. Our major aim is to clarify the health-promoting effects of fermented foods from the view point of microbiology. The filamentous fungus A. oryzae is a key microbe in the production of various fermented foods as it produces hydrolyzing enzymes to degrades raw materials such as grains and soy bean. Our recent studies have revealed that A. oryzae produces significant amount of agmatine, a natural polyamine that improves the quality of life (QOL), highlighting the significance of A. oryzae not only in the source of hydrolyzing enzymes but also in the enhancement of our health. We are investigating the molecular mechanisms of the physiologically active polyamine production by A. oryzae . The resultant findings are expected applicable for the development of novel fermented foods and nutraceuticals with increased functionalities to promote healthy life span in aging/aged society (Fig. 3).
References
- Akasaka et al., Amino Acids, 58, 13, 2026
- Yoshioka et al., Appl. Environ. Microbiol., 91, e01730-25, 2025
- Akasaka et al., J. Biotechnol., 405, 159-168, 2025
- Murakami et al. Appl. Environ. Microbiol., 90, e0029424, 2024
- Watanabe, Biosci. Biotechnol. Biochem., 88, 237, 2024
- Watanabe et al., Int. J. Mol. Sci., 25, 304, 2024
- Watanabe et al., NPJ Sci. Food, 7, 37, 2023
- Watanabe and Hashimoto, Sci. Rep., 13, 9279, 2023
- Ishii et al., J. Bacteriol., 203, e0016221, 2021
 NAIST Edge BIO
NAIST Edge BIO