Plant Metabolic Regulation

- Professor
- DEMURA Taku

- Assistant Professor
- MIZUTANI Miya

- Labs HP
- https://bsw3.naist.jp/demura/
Outline of Research and Education
Our laboratory engages in research and education pertaining to the biotechnology needed to resolve the issues facing human beings in the 21st century, such as food, environment, and energy. Especially we are exploring the mechanisms of gene expression regulation for woody cell differentiation using omics technology to develop novel biotechnological tools for the establishment of a sustainable society.
Major Research Topics
Molecular mechanisms governing xylem cell differentiation
Xylem functions in conduction of water and minerals throughout the plants, and supports the plant body. One of the features of xylem cells is development of secondary wall structure between plasma membrane and (primary) cell wall. Since woody biomass derived from xylem cells, xylem vessels and fiber cells, is one of important resources of land plant biomass, modifications of molecular mechanisms for xylem cell differentiation should be important strategies to improve plant biomass resources.
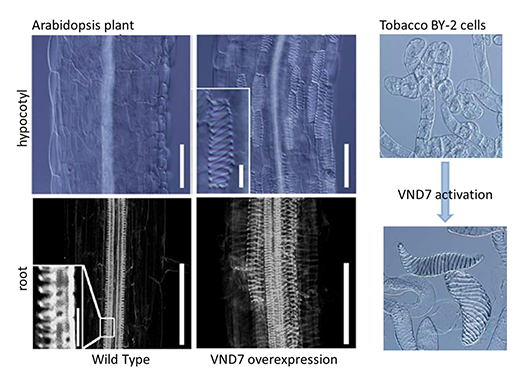
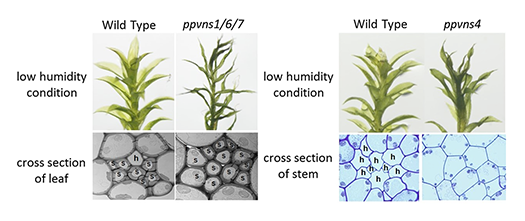
We identified a key regulator of the xylem vessel differentiation, Arabidopsis VND7 (VASCULAR-RELATED NAC-DOMAIN7), which is a plant-specific NAC domain transcription factor (Fig. 1). To understand the molecular mechanism by which xylem vessel formation is regulated, we have been characterizing VND7 and its homologs through various approaches. Recently we investigated the VND-homologous genes of moss Physcomitrella patens, and found that the VND-homologous genes function in the differentiation of water-conducting and supporting cells in the moss (Fig. 2). The VND-based molecular system is thus conserved among current land plants widely, suggesting that our findings can be applied to wide-range land plants to modify woody biomass.
We are also conducting genomics, transcriptome, proteome and metabolome studies to reveal the molecular system of plant biomass biosynthesis, using not only model plants but also non-model practical plants. Based on the obtained information, the designed transgenic poplars were generated and tested for the useful trains, such as higher stress tolerance, plant growth, and woody biomass accumulation (Fig. 3). These studies will give us the insights into effective biotechnological strategy to improve the quality and quantity of woody biomass.
Mechanism of plant mechanical optimization system
Plants change their structures into mechanically optimal shapes in the process of development and environmental response. We are analyzing the mechanical optimization system of plants on various scales (molecule-cell-tissue-organ) and elucidating its mechanism (Fig 4). We are performing the mechanical analysis using atomic force microscope (AFM), the 3D/4D structural analysis using time-lapse camera, micro X-ray CT and IoT devices (Raspberry Pi, etc.), and these analyses combined with the mathematical analysis. Based on the research findings, we are aiming to develop the application for establishing next-generation bio-basic technologies such as sophistication of plant function, and for creating sustainable architecture that is in harmony with various environmental factors such as earthquakes, typhoons, and temperature.
Development of Core Technologies for the Social Implementation of Autoluminescent Plants
Autoluminescent organisms such as bioluminescent mushrooms possess intrinsic light emitting systems. By introducing these bioluminescence pathways into plants through genetic engineering, it is possible to generate plants capable of autonomous light emission. In collaboration with Professor Takeharu Nagai at Osaka University, our laboratory has successfully produced autoluminescent poplar (Fig. 5). For future social implementation—such as utilizing autoluminescent plants as a source of illumination—further functional optimization, including enhancement of luminescence intensity, will be essential. We are already pursuing strategies to increase light output, including chemical induction methods and approaches that restrict autoluminescence to specific developmental stages or particular cell types. Through these efforts, we aim to advance the practical deployment of autoluminescent plants in society.

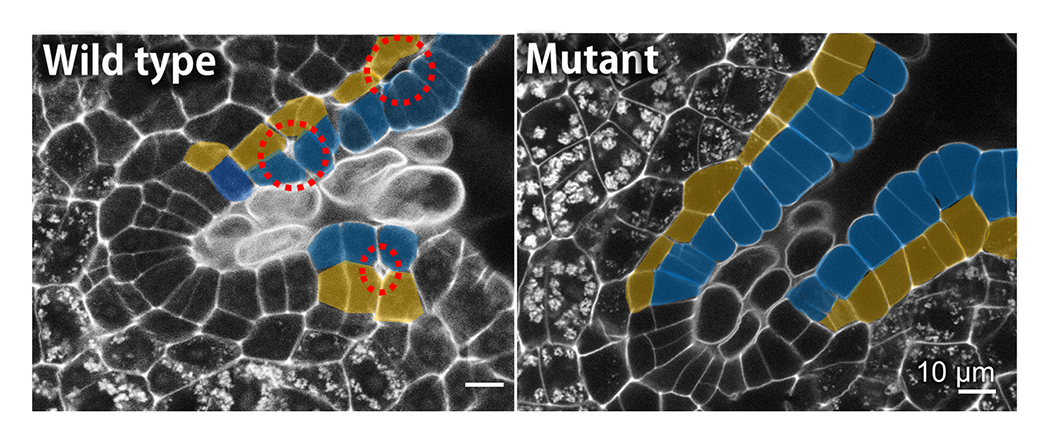
Intercellular spaces can be formed through the separation of adjacent cells, a process that involves the degradation and subsequent re-synthesis of cell wall components. Using M. polymorpha as a model system, we aim to elucidate the molecular mechanisms underlying this phenomenon.
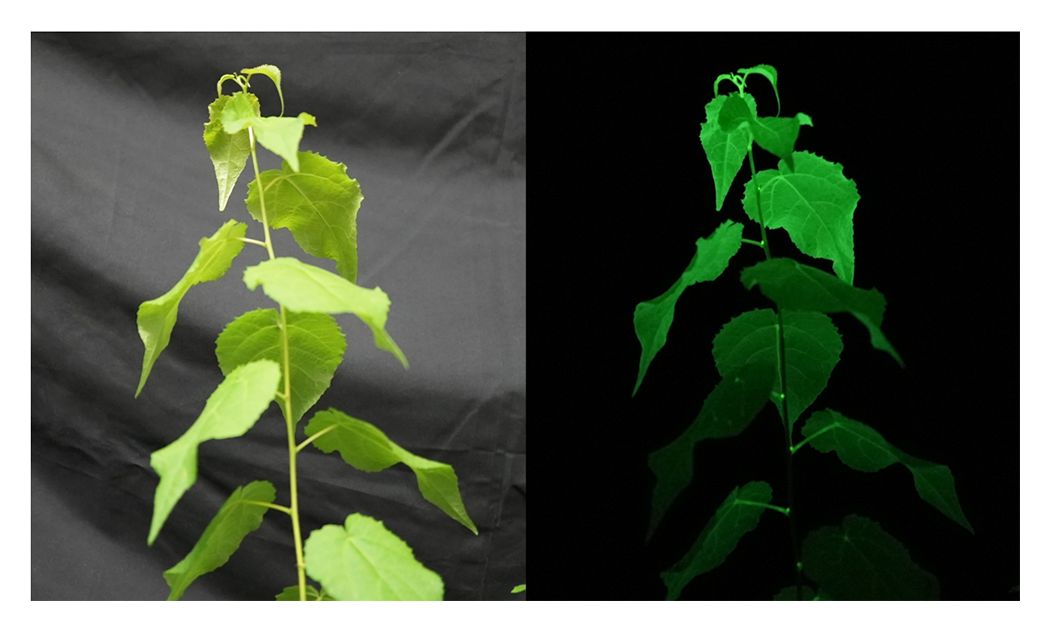

By introducing the bioluminescence system found in light emitting organisms such as bioluminescent mushrooms into poplar, we successfully generated autoluminescent poplar. We aim to advance the social implementation of such autoluminescent plants.
References
- Miyake A. et al., Curr. Biol., 36, 1086-1095, 2026
- Koyama T. et al., New Phytol., 248, 2981-2995, 2025
- Kunieda T. et al., Electronics Comm. Japan, 107, e12448, 2024
- Phookaew P. et al., Plant Cell, 36, 3298-3317, 2024
- Takahara M. et al., Plant Physiol., 192, 857-870, 2023
- Nakano Y. et al., Front. Plant Sci., 13, 819360, 2022
- Hirai R. et al., Front. Plant Sci., 13, 825810, 2022
- Akiyoshi N. et al., Plant Cell Physiol., 62, 1963-1974, 2021
- Terada et al., Plant Mol. Biol., 106, 309-317, 2021
- Nakata M.T. et al., Front. Plant Sci., 12, p654655, 2021
- Ramachandran V. et al., Plant Mol Biol, 104, 263-281, 2020
- Yoneda A. et al., Plants, 9, 604, 2020
- Tsugawa S. et al., Plant Biotechnol, 37, 443-450, 2020
- Kunieda T. et al., Plant Biotechnol, 37, 465-469, 2020
- Nakata M.T. et al., Plant Biotechnol, 37, 471-474, 2020
- Kunieda T. et al., Plant Cell Physiol., 61, 308-317, 2020
- Akiyoshi N. et al., Tree Physiol., 40, 704-716, 2020
- Tamura T. et al., Plant J., 100, 298-313, 2019
- Kubo M. et al., Nucleic Acids Res., 47, 4539-4553, 2019
- Ohtani M. et al., Curr. Opin. Biotech, 56, 82-87, 2019
- Takenaka Y. et al., Plant Cell, 30, 2663-2676, 2018
- Noguchi M. et al., Plant Biotechnol, 35, 31-37, 2018
- Tan T. et al., Plant Physiol, 176, 773-789, 2018
- Xu B. et al., Science, 343, 1505-1508, 2014
 NAIST Edge BIO
NAIST Edge BIO