Multicellular Network

- Professor
- SASAI Noriaki

- Assistant Professor
- SHINOZUKA Takuma

- MINEGISHI Takunori

- Labs HP
- https://bsw3.naist.jp/sasai/
Outline of Research and Education
Decoding the nervous system, advancing next-generation therapies
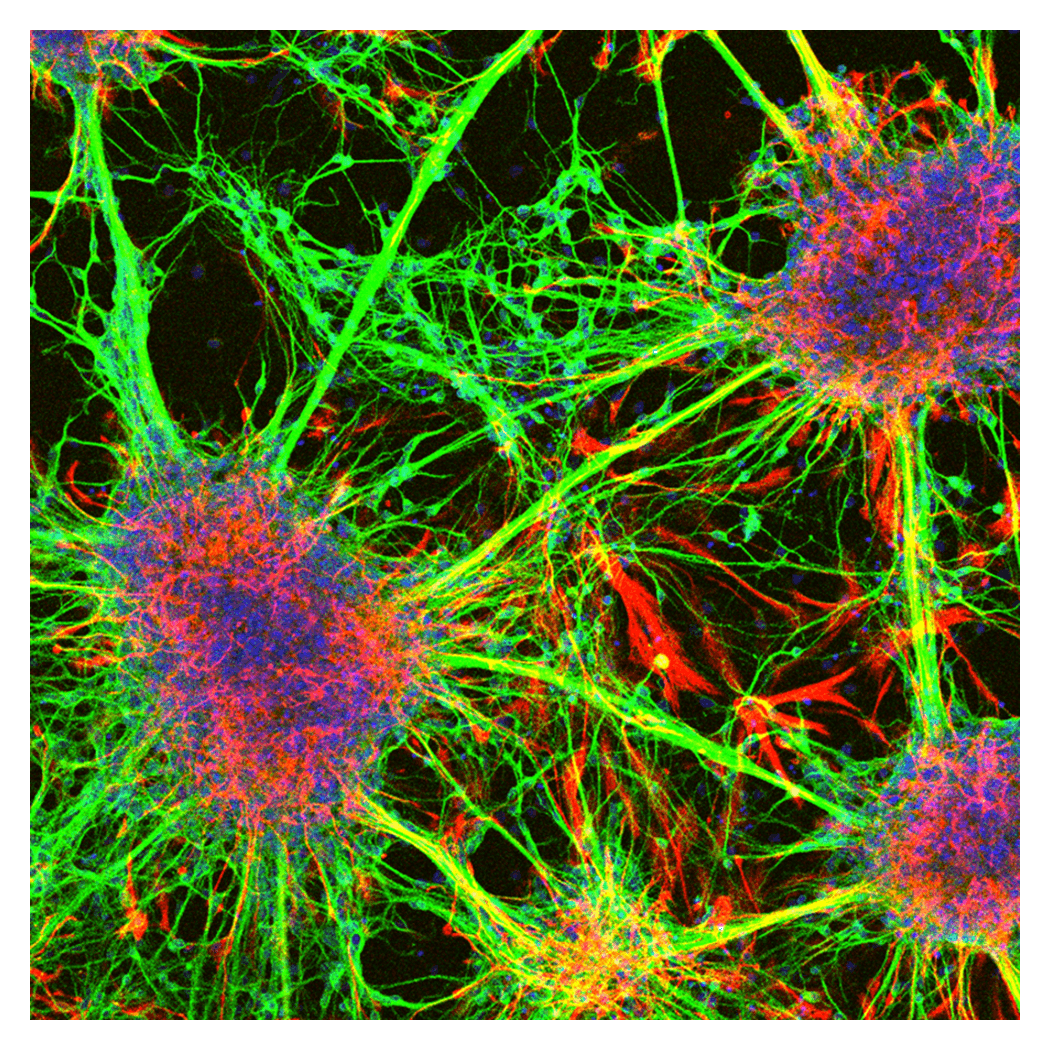
The central nervous system is the organ that controls an individual’s sensations and behavior, and it is composed of many different types of cells. These cells are formed mainly during embryonic development, and their arrangement and numbers are tightly regulated. To clarify these mechanisms, we conduct research using chick and mouse embryos, as well as stem cells such as ES cells and iPS cells, as experimental systems.
Because the nervous system has little capacity for natural recovery once damaged, its function must be maintained throughout life in order to preserve the organism’s abilities. With this in mind, we focus on hereditary eye diseases as examples of disrupted homeostasis. By elucidating the mechanisms underlying their onset, we aim to reveal how the functions of neural cells are maintained.
In this way, while pursuing fundamental research in the natural sciences centered on developmental biology and cell biology, we also develop interdisciplinary research that applies these techniques and findings to the development of treatments for diseases.
Major Research Topics
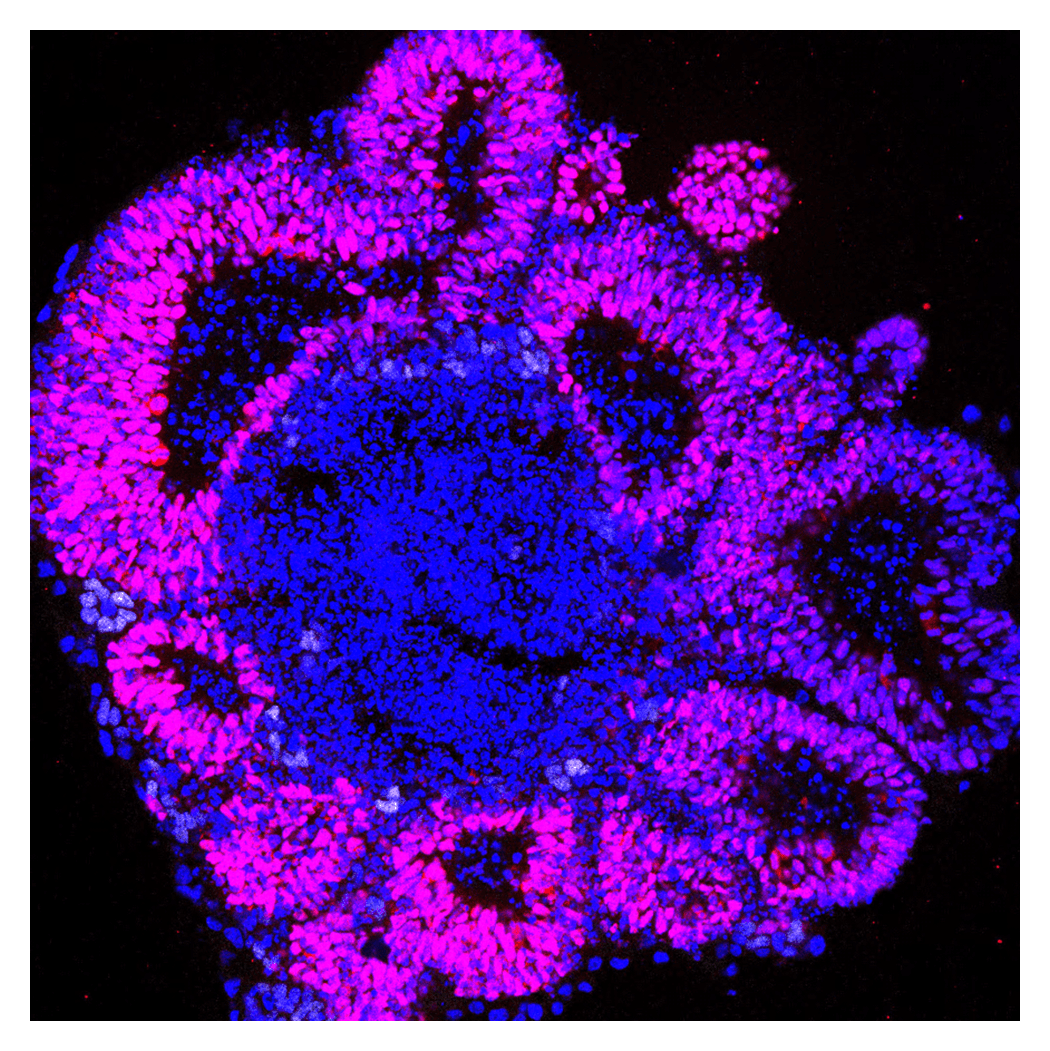
Decoding the mechanisms governing stem cell differentiation
During embryonic development, cells progress from an undifferentiated state through progenitor stages towards full maturation. This process is governed by a wide range of signalling molecules that play essential roles in cell proliferation and differentiation. We aim to elucidate the molecular basis of cell fate determination by investigating not only the functions of these signalling pathways, but also the underlying gene regulatory networks and dynamic changes in chromatin state that drive cellular differentiation.
Balancing proliferation and differentiation in tissue formation
Tissue size and cell number remain remarkably consistent across individuals. Such robust control of tissue size is thought to depend on a precise balance between cell proliferation and differentiation. We seek to define the genetic programmes that govern this balance and to reveal how they ensure the proper formation and maintenance of tissue architecture.
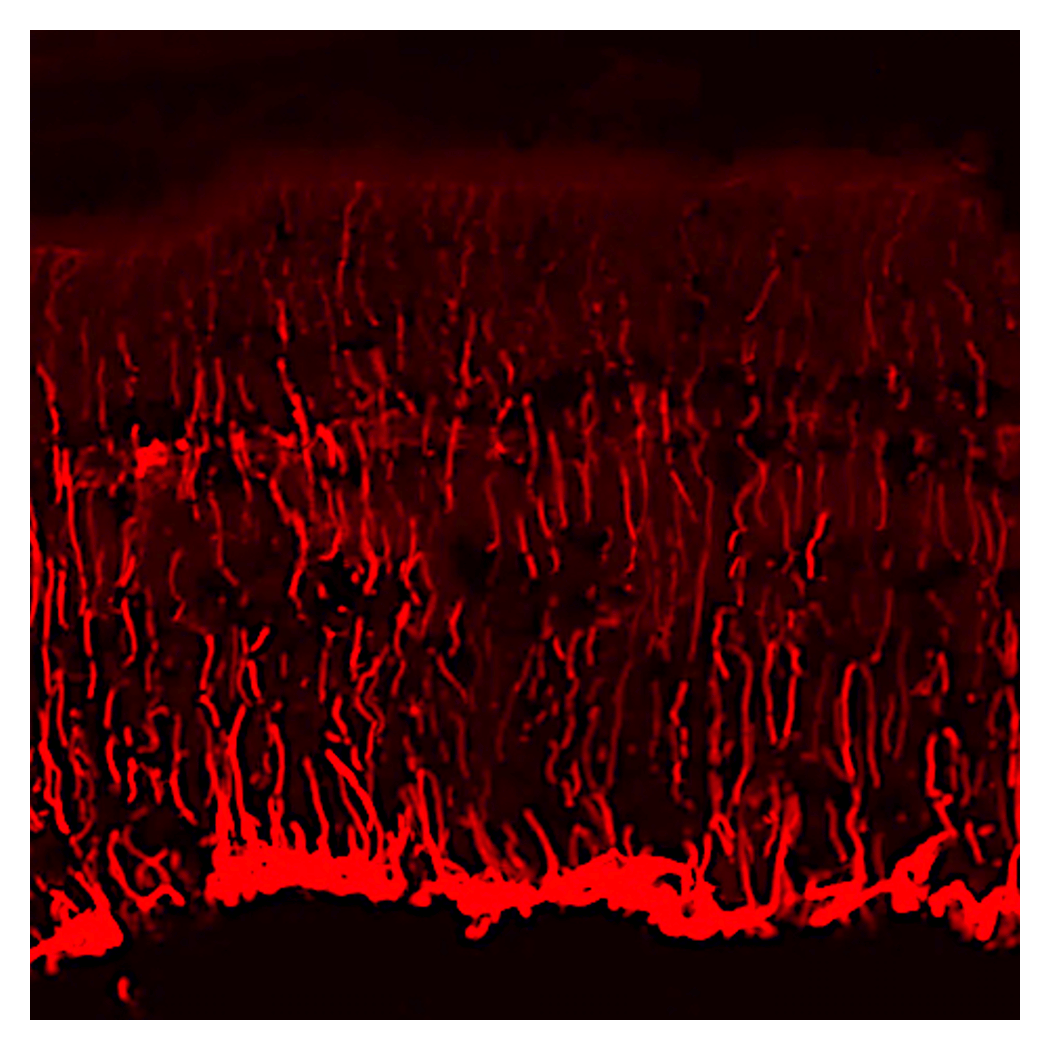
Pioneering novel therapies for inherited retinal diseases
Neural tissues, including the visual system, have limited regenerative capacity and rarely recover once damaged. As a result, many neurological disorders are progressive. Many are also rare diseases, making effective treatments difficult to develop. We focus on early gene expression changes shared across inherited retinal diseases. By applying this knowledge, we work to generate cells with greater resilience to external stress and lay the foundation for new treatments for blinding disorders.
References
- Matsumoto et al., Development Growth and Differentiation, 2025
- Shigesada et al., Cell Mol. Life Sci., 81, 51, 2024
- Sasai et al., Development Growth and Differentiation, 66, 89, 2024
- Ong et al., iScience, 26, 107887, 2023
- Yamamoto et al., Stem Cells, 41, 453, 2023
- Yamamoto#, Ong# et al., Development Growth and Differentiation, 64, 318, 2022
- Katsuyama et al., Developmental Dynamics, 251, 350-361, 2022
- Kobayashi et al., Disease Models and Mechanisms, 144, dmm048962, 2021
- Yatsuzuka et al., Development, 146, dev176784, 2019
- Kadoya and Sasai, Frontiers in Neuroscience, 13, 1022, 2019
- Hori et al., Sci. Rep. 9, 15911, 2019
 NAIST Edge BIO(
NAIST Edge BIO(