Plant Physiology

- Professor
- ENDO Motomu

- Assistant Professor
- KUBOTA Akane

- TAKAHASHI Nozomu

- Labs HP
- https://bsw3.naist.jp/endo/
Outline of Research and Education
Most living organisms possess internal mechanisms to measure time and seasons, using them to predict and respond to periodic environmental changes in advance.
In our laboratory, we aim to elucidate how plants measure time and seasons and their underlying molecular mechanisms, with a primary focus on the circadian clock, which ticks in an approximate 24-hour rhythm. Would you like to join us in challenging the mysteries of "time," which lie at the foundation of diverse plant physiological responses, such as gene expression, cell elongation, and photoperiodic flowering? Furthermore, our laboratory offers opportunities to engage in research from a bioinformatics perspective. We welcome approaches that seek to uncover the principles of the circadian clock not only through biological experimental methods but also through large-scale data analysis and the development of novel detection methods.
Through activities in our laboratory, you will not only gain a deeper understanding of plant physiology but also acquire logical thinking skills—the ability to identify the core of a problem and establish appropriate verification methods—which will serve as a lifelong asset for researchers and professionals.
Major Research Topics
Elucidating the mechanisms of flowering at specific timings: From seasonal to multi-decade scales
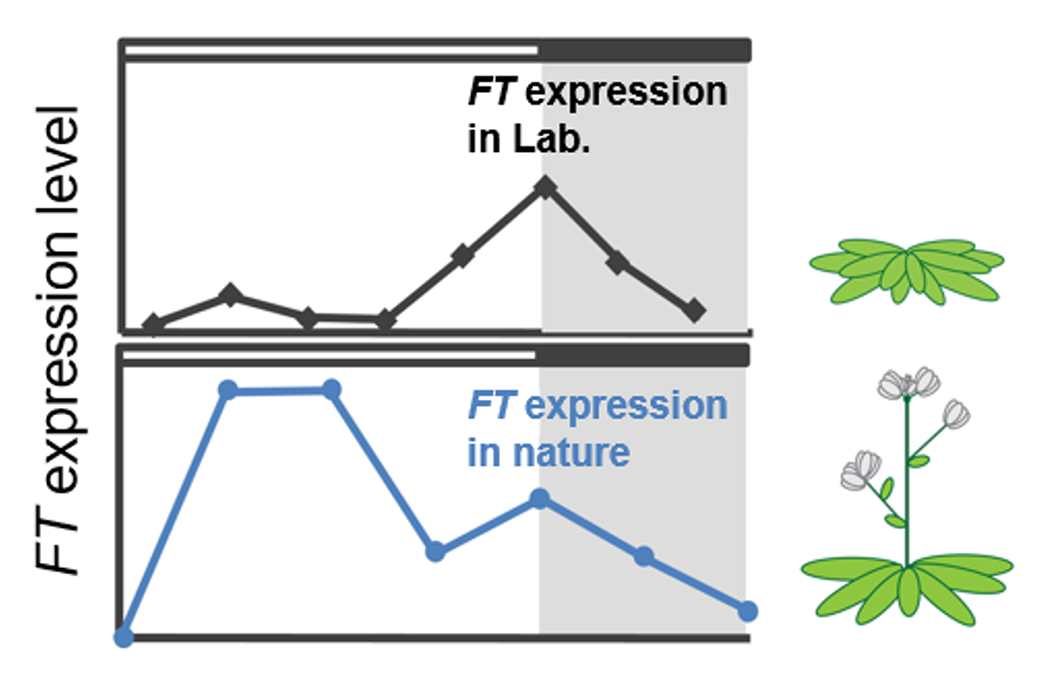
Seasonal photoperiodic flowering is controlled by the circadian clock. We have adopted a cutting-edge approach that goes beyond conventional laboratory conditions (constant temperature, no far-red light) to analyze plants under field conditions that include temperature fluctuations and far-red light. In the model plant Arabidopsis thaliana grown under field environments, it was revealed that the expression of the FT gene, which encodes the flowering hormone (florigen), has a distinct peak not only in the evening but also in the morning, leading to earlier flowering timing (Fig.1) Focusing on this previously overlooked morning peak of FT expression and the influence of temperature, we are uncovering unknown molecular mechanisms by integrating genetic, biochemical, and ecological perspectives.
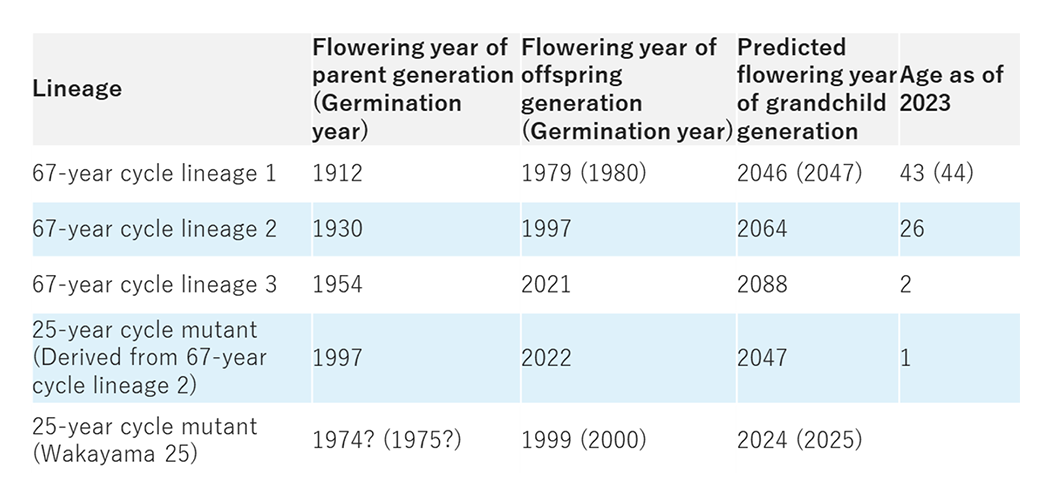
Additionally, elucidating the mechanism for measuring periods longer than a single day is a highly original theme. For instance, using Moso bamboo (three lineages exist in Japan), which is recorded to flower on a 67-year cycle, and a 25-year cycle lineage thought to have a mutation in its 'time-counting' mechanism, we are challenging the grand-scale mystery of how plants measure long-term periods spanning decades. We approach this through comprehensive genomic analysis using next-generation sequencers, tracking epigenetic modifications, and analyzing cultured bamboo cells (Fig.2)
Mechanisms and significance of long-distance transmission of time information
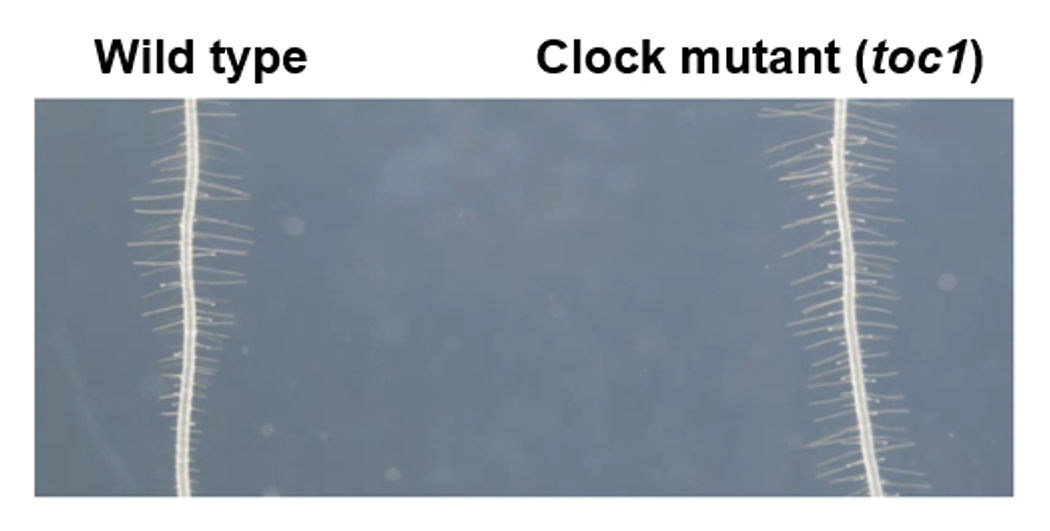
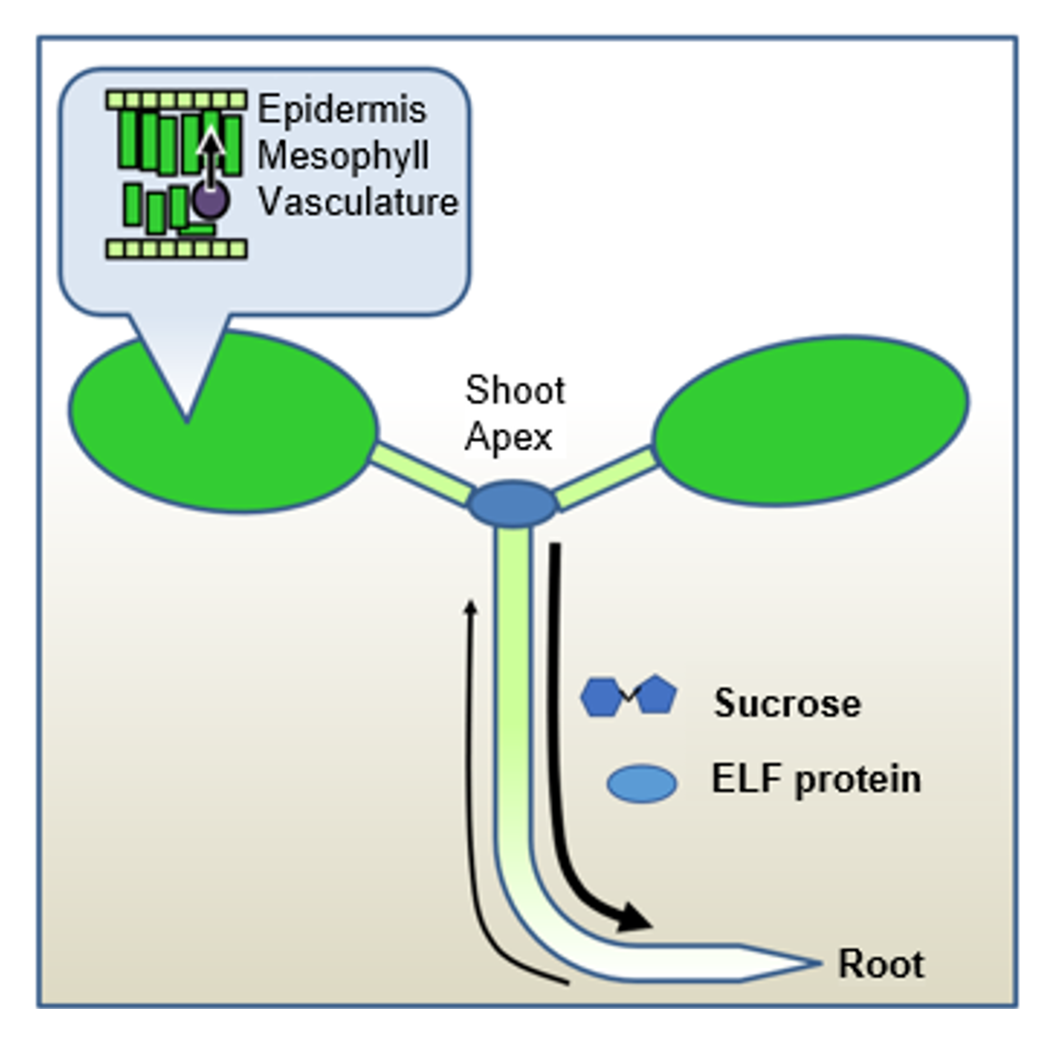
Although plants lack nerves or blood vessels, they must share the time measured in various parts of their bodies with other organs and tissues to grow as an integrated organism. Utilizing advanced reporter analysis employing grafting techniques and the firefly luciferase system, we have clarified the characteristics of the circadian clock at the organ and tissue levels. We are currently investigating how time information is exchanged between shoots (shoot apex) and roots, or between the vascular bundles and mesophyll of leaves, with a focus on sugars like sucrose, the mobile clock protein ELF4, and various nutrients (Fig.3). Furthermore, we recently discovered that a circadian rhythm also exists in the length of root hairs. It is known that signals transmitted from the shoots are crucial for this periodic elongation. To elucidate the true nature of these signals and the complete regulatory network, we are analyzing the significance and mechanisms of long-distance time information transmission, utilizing techniques such as grafting and automated quantification via deep learning (Fig.4).
Comprehensive analysis of oscillating genes using bioinformatics
The oscillating rhythm of gene expression is the foundation for generating periodicity in biological phenomena, yet there is still no unified consensus on the extent of oscillating genes throughout the plant. Our laboratory conducts analyses not only through experimental approaches but also from a bioinformatics perspective. We manage a time-series database of plant gene expression named GEAR (Gene Expression Archive for Rhythms), providing foundational data for rhythm researchers worldwide. Furthermore, using this massive time-series data, we are developing new computational methods to detect oscillating plant genes more accurately. By comprehensively identifying rhythmic genes that were previously overlooked as noise by conventional methods, we aim to uncover the fundamental principles of rhythm generation.

References
- Endo et al. Nucleic acids research, 54(D1), D1711–D1719, 2026
- Ikeda et al., Plant Cell Physiol, 64,1289-1300, 2023
- Torii K, et al., Cell Reports, 61, 243-254, 2020
- Chen, Takahashi et al., Nat Plants., 6, 416-425, 2020
- Song, Kubota et al., Nat Plants., 4, 824-835, 2018
- Endo et al., Nat Protoc., 11, 1388-1395, 2016
- Shimizu et al., Nat Plants., 1, 15163, 2015
 NAIST Edge BIO(
NAIST Edge BIO( 