Plant Symbiosis

- Professor
- YOSHIDA Satoko

- Assistant Professor
- WAKATAKE Takanori

- HIRATA Rikako

- Labs HP
- https://bsw3.naist.jp/yoshida/index-en.html
Outline of Research and Education
Parasitic plants - major agricultural constrains in the world
Parasitic plants are able to parasitize other plants and rely on their hosts for water and nutrients. Several parasitic plants in the Orobanchaceae family, such as Striga (Fig. 1) and Orobanche spp., cause enormous damage to world agriculture because they parasitize important crops and vegetables. We are investigating molecular mechanisms underlying plant parasitism using the model parasitic plants Phtheirospermum japonicum and weedy parasite Striga spp. By combining molecular, genetic, cell biology and genomic approaches, we aim to understand the nature of parasitism and eventually develop novel control methods for weedy parasites.
Major Research Topics
Identification of genes involved in haustorium formation
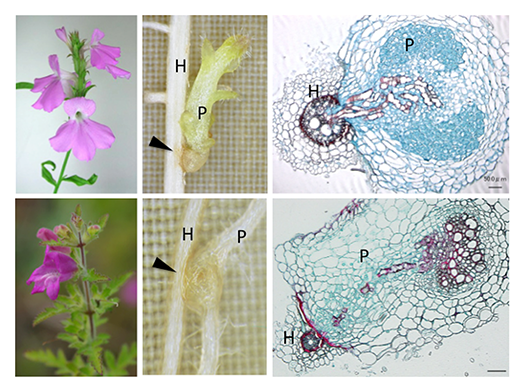
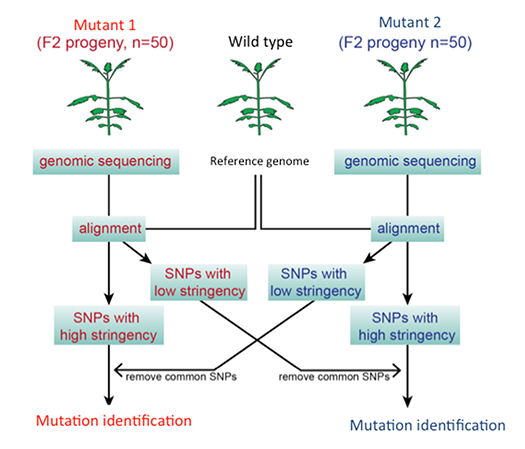
Parasitic plants form specialized invasive organs called “haustorium”. The haustorium invades host roots, and eventually forms a vasculature connection between the host and the parasite to assimilate host nutrients (Fig. 2). To identify the genes involved in haustorium formation, forward and reverse genetic tools in P. japonicum were established. Screening of P. japonicum mutants which lack haustorium formation and identification of the causal genes by next-generation sequencing (Fig. 3) will isolate the essential genes in the haustorium formation. Furthermore, the genes upregulated during haustorium formation will be reverse-genetically analyzed.
Plant-plant communication via small-molecular weight compounds
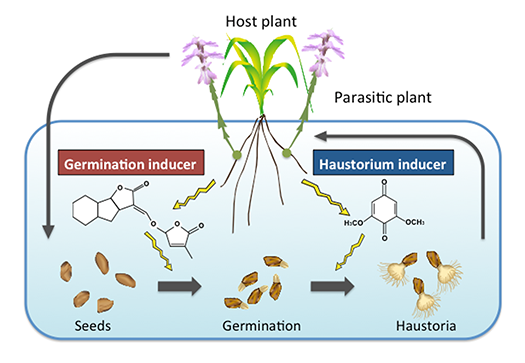
Parasitic plants recognize their hosts via small-molecular weight compounds secreted from the host plant (Fig. 4). For example, the obligate parasite Striga germinates in response to the plant hormone strigolactones. The haustorium formation is induced by derivatives of cell wall lignin; however, the nature of haustorium inducers has not been clearly understood. We are trying to identify novel haustorium inducing compounds.
Comparative genomics of parasitic plants
Recent progress in next-generation sequencing technology enables us to acquire the complete genome sequence of any plant. We sequenced the whole genomes of Striga and P. japonicum. By examining these genome sequences, we found that parasitic plants have experienced evolutional events such as expansion of specific gene family and horizontal gene transfers from hosts. How did the plants obtain new genes, increase the copy numbers and eventually acquire a new trait? What is the genetic diversity among Striga species in Africa? We analyze genome evolution using bioinformatics tools.

References
- Suga et al., Plant Physiol., 200, kiaf686, 2026
- Xiang et al. Science, 390, 405-410, 2025
- Aoki et al. Plant Cell Physiol., 63, 1446-1456, 2022
- Masumoto et al., Plant Physiol., 185, 1429-1442, 2021
- Yoshida and Kee, Curr Opin Biotech, 70,248-254, 2021
- Furuta et al., Plant Physiol., 186, 1424-1434,2021
- Cui et al., Science Advances, 6, eabc2385, 2020
- Yoshida, S. et al., Curr. Biol., 18, 3041-3052, 2019
- Cui, S. et al., New Phytologist, 218, 710-723, 2018
- Yoshida, S. et al., Ann. Rev. Plant Biol., 67, 643-67, 2016
 NAIST Edge BIO(
NAIST Edge BIO( 