Research Projects
1. Novel stress-tolerant mechanisms of yeast cells and their applications for breeding of industrial yeast
5) Development of industrial yeast based on novel stress-tolerant mechanisms
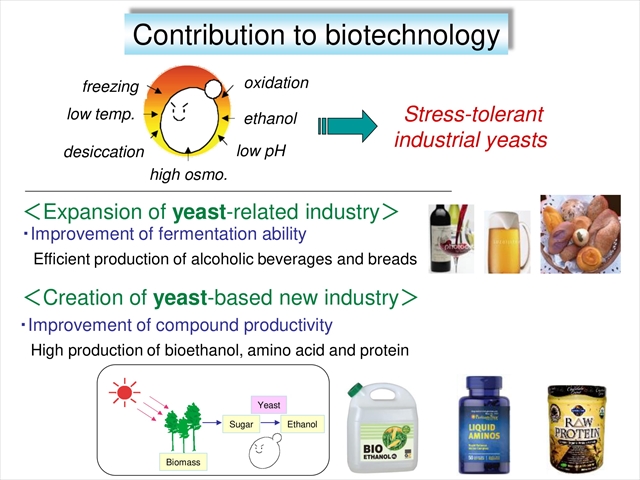
Through our basic research on the novel stress-tolerant mechanisms, we will construct industrial yeasts with higher tolerance to various stresses and contribute to yeast-based industry for the effective production of frozen dough and alcoholic beverages or the breakthrough of bioethanol production.
- H. T. Phuong, Y. Ishiwata-Kimata, Y. Nishi, N. Oguchi, H. Takagi and Y. Kimata: Aeration mitigates endoplasmic reticulum stress even without mitochondrial respiration in Saccharomyces cerevisiae. Microbial Cell, 8, 77-86 (2021).
- H. Takagi: Molecular mechanisms and highly-functional development for stress tolerance of the yeast Saccharomyces cerevisiae. Biosci. Biotech. Biochem., 85, 1017-1037 (2021). DOI: 10.1093/bbb/zbab022.
- Y. Nabilah Binti Mohd Fauzee, N. Taniguchi, Y. Ishiwata-Kimata, H. Takagi and Y. Kimata: The unfolded protein response in Pichia pastoris without external stressing stimuli. FEMS Yeast Res., 20, foaa053, in press. DOI: 10.1093/femsyr/foaa053.
- M. E. Prastya, R. I. Astuti, I. Batubara, H. Takagi and A. T. Wahyudi: Natural extract and its fractions isolated from the marine bacterium Pseudoalteromonas flavipulchra STILL-33 have antioxidant and antiaging activities in Schizosaccharomyces pombe. FEMS Yeast Res., 20, foaa014 (2020). DOI: 10.1093/femsyr/foaa014.
- M. E. Prastya, R. I. Astuti, I. Batubara, H. Takagi and A. T. Wahyudi: Chemical screening identifies an extract from marine Pseudomonas sp.-PTR-08 as an anti-aging agent that promotes fission yeast longevity by modulating the Pap1-ctt1+ pathway and the cell cycle. Mol. Biol. Rep., 47, 33-43 (2020).
- D. Watanabe, S. Tashiro, D. Shintani, Y. Sugimoto, A. Iwami, Y. Kajiwara, H. Takashita and H. Takagi: Loss of Rim15p in shochu yeast alters carbon utilization during barley shochu fermentation. Biosci. Biotech. Biochem., 83, 1594-1597 (2019).
- D. Watanabe, T. Kajihara, Y. Sugimoto, K. Takagi, M. Mizuno, Y. Zhou, J. Chen, K. Takeda, H. Tatebe, K. Shiozaki, N. Nakazawa, S. Izawa, T. Akao, H. Shimoi, T. Maeda and H. Takagi: Nutrient signaling via the TORC1-Greatwall-PP2AB55δ pathway responsible for the high Initial rates of alcoholic fermentation in sake yeast strains of Saccharomyces cerevisiae. Appl. Environ. Microbiol., 85(1), e02083-18 (2019).
- D. M. Tran, H. Takagi and Y. Kimata*: Categorization of endoplasmic reticulum stress as accumulation of unfolded proteins or membrane lipid aberrancy using yeast Ire1 mutants. Biosci. Biotech. Biochem., 83, 326-329 (2019).
- T. C. Mai, T. Munakata, D. M. Tran, H. Takagi and Y. Kimata*: A chimeric mutant analysis in yeast cells suggests BiP independent regulation of the mammalian endoplasmic reticulum-stress sensor IRE1α. Biosci. Biotech. Biochem., 82, 1527-1530 (2018).
- M. Oomuro, D. Watanabe, Y. Sugimoto, T. Kato, Y. Motoyama, T. Watanabe and H. Takagi: Accumulation of intracellular S-adenosylmethionine increases the fermentation rate of bottom-fermenting brewer's yeast during high-gravity brewing. J. Biosci. Bioeng., 126, 736-741 (2018).
- D. Watanabe, M. Kumano, Y. Sugimoto, M. Ito, M. Ohashi, K. Sunada, T. Takahashi, T. Yamada and H. Takagi: Metabolic switching of sake yeast by kimoto lactic acid bacteria through the [GAR+] non-genetic element. J. Biosci. Bioeng., 126, 624-629 (2018).
- D. A. Nur’utami, L. Haditjaroko, H. Takagi and K. Syamsu: Hyperosmotic stress tolerance of transcription activator Msn2-Over expression strain and proline-NO synthesis strain of Saccharomyces cerevisiae in very high gravity bioethanol fermentation. Pak. J. Biotechnol., 14, 135-139 (2017).
- D. Watanabe, H. Takagi: Pleiotropic functions of the yeast Greatwall-family protein kinase Rim15p: a novel target for the control of alcoholic fermentation. Biosci. Biotech. Biochem., 81, 1061-1068 (2017).
- D. Watanabe, A. Kaneko, Y. Sugimoto, T. Negishi, S. Ohnuki, H. Takagi and Y. Ohya: Promoter engineering of the Saccharomyces cerevisiae RIM15 gene for improvement of alcoholic fermentation rates under stress conditions. J. Biosci. Bioeng., 123, 183-189 (2017).
- D. Watanabe, Y. Zhou, A. Hirata, Y. Sugimoto, K. Takagi, T. Akao, Y. Ohya, H. Takagi and H. Shimoi*: Inhibitory role of Greatwall-like protein kinase Rim15p in alcoholic fermentation via upregulating the UDP-glucose synthesis pathway in Saccharomyces cerevisiae. Appl. Environ. Microbiol., 82, 340-351 (2016).
- S. Hirayama, M. Shimizu, N. Tsuchiya, S. Furukawa, D. Watanabe, H. Shimoi, H. Takagi, H. Ogihara and Y. Morinaga: Awa1p on the cell surface of sake yeast inhibits biofilm formation and the co-aggregation between sake yeasts and Lactobacillus plantarum ML11-11. J. Biosci. Bioeng., 119, 532-537 (2015).
- T. Inaba, D. Watanabe, Y. Yoshiyama, K. Tanaka, J. Ogawa, H. Takagi, H. Shimoi and J. Shima: An organic acid-tolerant HAA1-overexpression mutant of an industrial bioethanol strain of Saccharomyces cerevisiae and its application to the production of bioethanol from sugarcane molasses. AMB Express, 3:74, doi:10.1186/2191-0855-3-74 (2013).
- H. Kitagaki and H. Takagi: Mitochondrial metabolism and stress response of yeast: Applications in fermentation technologies. J. Biosci. Bioeng., 117, 383-393 (2013).
- Y. Sasano, Y. Haitani, K. Hashida, S. Oshiro, J. Shima and H. Takagi: Improvement of fermentation ability under baking-associated stress conditions by altering the POG1 gene expression in baker's yeast. Int. J. Food Microbiol., 165, 241-245 (2013).
- T. Inai, D. Watanabe, Y. Zhou, R. Fukada, T. Akao, J. Shima, H. Takagi and H. Shimoi: Rim15p-mediated regulation of sucrose utilization during molasses fermentation using Saccharomyces cerevisiae strain PE-2. J. Biosci. Bioeng., 116, 591-594 (2013).
- S. Hasegawa, T. Ogata, K. Tanaka, A. Ando, H. Takagi and J. Shima: Overexpression of vacuolar H+-ATPase-related genes in bottom-fermenting yeast enhances ethanol tolerance and fermentation rates during high-gravity fermentation. J. Inst. Brew., 118, 179-185 (2012).
- Y. Sasano*, Y. Haitani*, K. Hashida, I. Ohtsu, J. Shima and H. Takagi: Overexpression of the transcription activator Msn2 enhances fermentation ability of industrial baker’s yeast in frozen dough. *These authors contributed equally to this work. Biosci. Biotech. Biochem., 76, 624-627 (2012).
- Y. Sasano*, D. Watanabe*, K. Ukibe, T. Inai, I. Ohtsu, H. Shimoi and H. Takagi: Overexpression of the yeast transcription activator Msn2 confers furfural resistance and increases the initial fermentation rate in ethanol production. *These authors contributed equally to this work. J. Biosci. Bioeng., 113, 451-455 (2012).
- H. Urbanczyk, C. Noguchi, H. Wu, D. Watanabe, T. Akao, H. Takagi, H. Shimoi: Sake yeast strains have difficulty in entering a quiescent state after cell growth cessation. J. Biosci. Bioeng., 112, 44-48 (2011).
- T. Nakamura, S. Takahashi, H. Takagi and J. Shima: Multicopy suppression of oxidant-sensitive eos1 mutation by IZH2 in Saccharomyces cerevisiae and the involvement of Eos1 in zinc homeostasis. FEMS Yeast Res., 10, 259-269 (2010).
- K. Ukibe, K. Hashida, N. Yoshida and H. Takagi: Metabolic engineering of Saccharomyces cerevisiae for astaxanthin production and oxidative stress tolerance. Appl. Environ. Microbiol., 75, 7205-7211 (2009).
- S. Takahashi*, A. Ando*, H. Takagi and J. Shima: Insufficiency of copper ion homeostasis causes freeze-thaw injury of yeast cells revealed by indirect gene expression analysis. *These authors contributed equally to this work. Appl. Environ. Microbiol., 75, 6706-6711 (2009).
- J. Shima and H. Takagi: Stress-tolerance of baker’s yeast (Saccharomyces cerevisiae) cells: stress-protective molecules and genes involved in stress tolerance. Biotechnol. Appl. Biochem., 53, 155-164, 2009.
- H. Wu, T. Watanabe, Y. Araki, H. Kitagaki, T. Akao, H. Takagi and H. Shimoi: Disruption of ubiquitin-related genes in laboratory yeast strains enhances ethanol production during sake brewing. J. Biosci. Bioeng., 107, 636-640, 2009.
- T. Nakamura, H. Takagi and J. Shima: Effects of ice-seeding temperature and intracellular trehalose contents on survival of frozen Saccharomyces cerevisiae cells. Cryobiol., 58, 170-174 (2009).
- Y. Araki, H. Wu, H. Kitagaki, T. Akao, H. Takagi and H. Shimoi: Ethanol stress stimulates the Ca2+-mediated calcineurin/Crz1 pathway in Saccharomyces cerevisiae. J. Biosci. Bioeng., 107, 1-6 (2009).
- T. Nakamura, S. Mizukami-Murata, A. Ando, Y. Murata, H. Takagi and J. Shima: Changes in gene expression of commercial baker's yeast during an air-drying process that simulates dried yeast production. J. Biosci. Bioeng., 106, 405-408 (2008).
- J. Shima, A. Ando and H. Takagi: Possible roles of vacuolar H+-ATPase and mitochondrial function in tolerance to air-drying stress revealed by genome-wide screening of Saccharomyces cerevisiae deletion strains. Yeast, 25, 179-190 (2008).
- F. Tanaka-Tsuno, S. Mizukami-Murata, Y. Murata, T. Nakamura, A. Ando, H. Takagi and J. Shima: Functional genomics of commercial baker’s yeasts that have different abilities for sugar utilization and high-sucrose tolerance under sugar conditions. Yeast, 24, 901-911 (2007).
- T. Nakamura, A. Ando, H. Takagi and J. Shima: Eos1, whose deletion confers sensitivity to oxidative stress, is involved in N-glycosylation in Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 353, 293-298 (2007).
- A. Ando, T. Nakamura, Y. Murata, H. Takagi and J. Shima: Identification and classification of genes required for tolerance to freeze-thaw stress revealed by genome-wide screening of Saccharomyces cerevisiae deletion strains. FEMS Yeast Res., 7, 244-253 (2007).
- H. Wu, X. Zheng, Y. Araki, H. Sahara, H. Takagi and H. Shimoi: Global gene expression analysis of yeast cells during sake brewing. Appl. Environ. Microbiol., 72, 7353-7358 (2006).
- F. Tanaka, A. Ando, T. Nakamura, H. Takagi and J. Shima: Functional genomic analysis of commercial baker’s yeast during initial stages of model dough-fermentation. Food Microbiology, 23, 717-728 (2006).
- M. Sugiura and H. Takagi: Yeast cell death caused by mutation of the OST2 gene encoding the -subunit of the Saccharomyces cerevisiae oligosaccharyltransferase. Biosci. Biotech. Biochem., 70, 1234-1241 (2006).
- A. Ando, F. Tanaka, Y. Murata, H. Takagi and J. Shima: Identification and classification of genes required for tolerance to high sucrose stress revealed by genome-wide screening of Saccharomyces cerevisiae. FEMS Yeast Research, 6, 249-267 (2006).
- M. Wada, S. Nakamori and H. Takagi: Serine racemase homologue of Saccharomyces cerevisiae has L-threo-3-hydroxyaspartate dehydratase activity. FEMS Microbiol. Lett., 225, 189-193 (2003).
- Y. Kubo, H. Takagi and S. Nakamori: Effect of gene disruption of succinate dehydrogenase on succinate production in a sake yeast strain. J. Biosci. Bioeng., 90, 619-624 (2000).
